Journal of HIV for Clinical and Scientific Research
Detection of katG Ser315Thr substitution in Mycobacterium tuberculosis infected patients using PCR -RFLP method for screening Isoniazid-Resistance
1Mapmygenome India Limited, Royal Demeure, Madhapur, Hyderabad -500081, India
2Lilac Insights Private Limited, Rupa Solitaire MIDC, Navi Mumbai-400710 India
Author and article information
Cite this as
Jadhav SK, Mote SR (2019) Detection of katG Ser315Thr substitution in Mycobacterium tuberculosis infected patients using PCR -RFLP method for screening Isoniazid-Resistance. J HIV Clin Sci Res. 2018; 6(1): 1-4. Available from: 10.17352/2455-3786.000128
Copyright License
© 2018 Jadhav SK, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.Currently mycobacterium tuberculosis is major health issue to the Indian population as well as global. According to WHO around 8 million cases of Tuberculosis occur each year, which resulting in approximately 3 million deaths and the situation is more complicated due to emergence of multidrug-resistant TB. It is the leading cause of death worldwide and results from a single infectious agent, ranking over HIV/AIDS.
Currently mycobacterium tuberculosis is major health issue to the Indian population as well as global. According to WHO around 8 million cases of Tuberculosis occur each year, which resulting in approximately 3 million deaths and the situation is more complicated due to emergence of multidrug-resistant TB. It is the leading cause of death worldwide and results from a single infectious agent, ranking over HIV/AIDS. In 2017, Tuberculosis caused an estimated of 1.3 million deaths among HIV-negative people and additional there were 300 000 deaths from Tuberculosis among people living with HIV. Globally around 10.0 million people which ranges from 9.0–11.1 million were developed Tuberculosis in 2017 which accounts 5.8 million men and 3.2 million women and along with 1.0 million were children’s. There were cases with all age groups reported worldwide but overall about 90% were adults (aged ≥15 years) and 9% were people living with HIV [1]. Rifampin and Isoniazid are two first line drugs involved in the treatment of TB when the resistantance was developed against these two drugs they are termed as multi drug resistant (MDR) Mycobacterium tuberculosis.
The rate of spread of MDR strain directly affects tuberculosis control program so for that rapid screening or identification of MDR strain creates major help in the treatment and control of MDR strains in the population. There is an approximate estimation of 79,000 multi-drug resistant Tuberculosis patients notified cases with pulmonary Tuberculosis each year. In 2016 an average estimate of 28 lakh cases were occurred and 4.5 lakh people were died due to Tuberculosis (TB facts). In drug resistant TB person will not respond to one of the main TB drug. Mostly in India the drug resistant TB is due to failure of people to take their anti TB drugs properly. According to WHO TB statistics for India provides an estimated incidence of 2.79 million cases. In most of the cases the multidrug resistant TB is resistant to rifampin and isoniazid drugs. In rifampin the resistant due to the mutation in rpoB gene region while isoniazid resistant involves genes like katG, inhA, kasA and aphC but most frequently it is associated with the katG mutation [2]. Mutation in the region of katG gene occurs in higher fold frequency than rpoB gene mutations and is considered as a step in the evolution of MDR-Tuberculosis [3]. Although Ser315Thr mutation is commonly found mutation used for identifying isoniazid resistant isolates [4].
Isoniazid inhibits the process of biosynthesis of cell wall with mycolic acid and thereby making the mycobacteria more susceptible to the reactive oxygen radicals and along with environmental factors [5]. The isoniazid also requires the catalytic activation to be in active form and Catalase-peroxidase is the key product of the katG gene. The mutation in this specific gene will cause the mycobacteria contributes for isoniazid drug resistantance. In this study we assess the PCR-RFLP technique to screen Ser315Thr substistution in katG region for isoniazid resistant Mycobacterium strains. In the treatment of tuberculosis Isoniazid is one most common first line drug if the resistant developed against this drug it will causes serious problems related to treatment. The most frequent mutation in katG gene is substitution from AGC to ACC at codon 315 [6]. The molecular method like sequencing is used for the detection of isoniazid resistant is time consuming therefore in the present study we evaluated the PCR-RFLP based rapid screening method. PCR-RFLP method is very quick for screening or detection of katG gene mutation in Isoniazid resistance Tuberculosis infected population.
Materials and Methods
A total of ten Sputum samples which are TB Confirmed positive samples were collected from primary health care centers Hyderabad. The patients are already confirmed with AFB and culture positive and these patients showed advance stage of immunological complications hence it is important to evaluate for multi drug resistance pattern for these patients for further effective therapeutic treatment. We have used the reference standards to evaluate the specificity and sensitivity of this assay. A result with reference and samples generated were compared with DNA sequencing result. The study was carried out to identify Ser315Thr mutation at specific position of katG gene for isoniazid resistant Mycobacterium tuberculosis strains.
Genomic DNA extraction
DNA extraction was carried out from the sputum samples using QiaAmp DNA Mini kit as per the manufacturer’s protocol. The purity of the DNA was quantified by A260/A280 ratio and the optical density for DNA was measured at 260 and 280 nm) using Nan drop ND-1000 spectrophotometer (Thermofisher Scientific) and the DNA was visualized on 0.8% agarose gel.
PCR-RFLP for katG
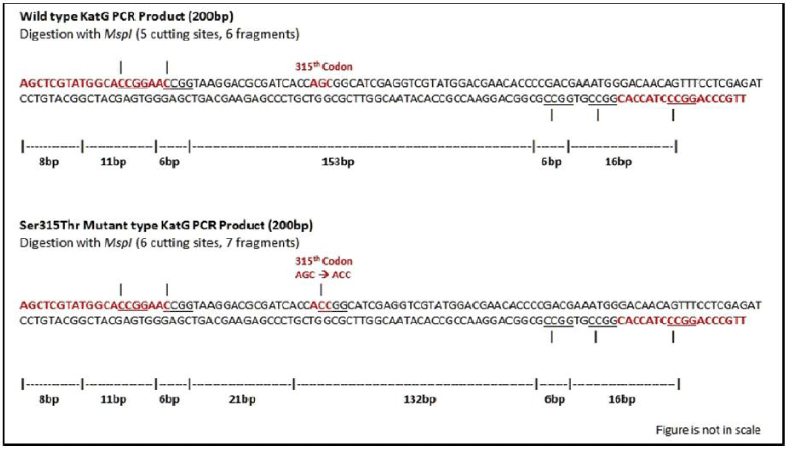
This PCR-RFLP method was designed to easily detect the katG specific codon mutation from AGC (Serine)→ ACC (Threonine) which leads Isoniazid resistance mycobacterium strains. ThePCR product of 200 bp fragment was amplified for identifying katG codon 315 was carried out with the specific primers katG F and katG R as reported [7], in a 25ul of a PCR mixture containing 0.4 picomoles of each primers and Taq polymerase 2X master mix (G-Bioscience in which Taq polymerase supplied with 2X Taq buffer, 0.4mM dNTPs, 3.2mM MgCl2and 0.02% bromophenol blue and the PCR was performed in thermal cycler with the following conditions, initial denaturation at 95˚C for 5min and 30 cycles of 94˚C for 1 min with 61°C for 1 min and 72˚C for 1 min and a final extension was carried out at 72˚C for 4 min. The reaction controls were used to detection false positive results due to contamination a negative control (distilled water) was included in each PCR reaction. A reference control H37Rv was used in each PCR run which shows a clear band at position of 200bp in the agarose gel electrophoresis (Table 1 and Figure 1).
Agarose gel electrophoresis for MTB detection
The PCR products were detected by 2% agarose gel electrophoresis with stained ethidium bromide. Mycobactrium strain H37Rv used as a reference control and negative control was also used in the PCR reaction.
Restriction enzyme digestion for Ser315Thr katG
The katG amplicons were further used for RFLP analysis using MspI digestion. To each of the PCR mixtures add restriction enzyme mixture containing a final concentration of 0.5U of MspI enzyme (NEB Biolabs), 1X NEBuffer used for reaction according to manufacturer’s instructions. The mixture incubated at 37°C for 1 hr. The restriction fragments were visualized by running on 2% agarose gel. This is very simple and rapid RFLP method used to detect the katG codon mutation from AGC to ACC, in which mutation creates an additional site for MspI (CCGG) and therefore it is cleaved by restriction enzyme. This mutation from AGC to ACC creates isoniazid drug resistant phenotype. As a result, 153 bp longest fragments indicates the wild type katG PCR product, while 132 bp long fragment indicates the presence of Ser315Thr katG mutant along with shorter bands sizes (21 bp-8 bp products) which cannot be resolved clearly on gel due to smaller fragment size. MTB strain H37Rv (Accession Number X68081) used as a reference control in the restriction digestion.
Results
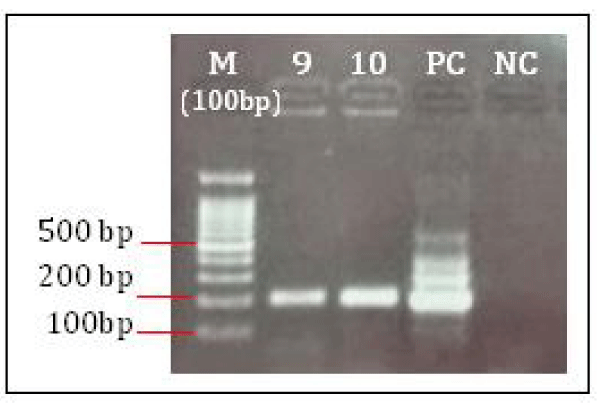
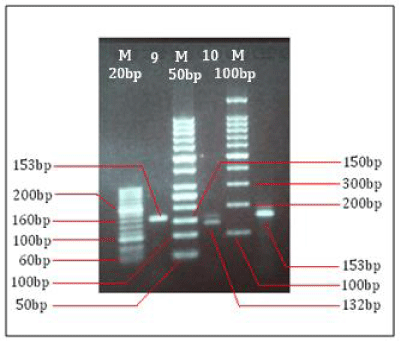
DNA extracted from ten isolates were visualized on 8% agarose gel displayed positive band (Figure 2). Further extracted DNA used for katG amplification, after amplification the amplified product were observed on 2% agarose gel for confirmation of 200bp positive amplification (Figure 3). After confirmation the katG amplicons were further used for RFLP analysis using MspI digestion. After digestion the electrophoresis of katG digested product shows out of 10 samples only one sample shows 132bp region results occurrence of Ser315Thr mutation while the other 9 samples shows 153bp region results no mutation. The (Figure. 4) shows that the products obtained by MspI digestion out of which lane 2 (sample 9) with undigested product or wild type; lane 4 (sample 10) with mutated katG (315ACC); last well is control; M represents the DNA ladder (G-Bioscience) lane 1 (20bp), lane 3 (50bp), lane 6 (100bp) respectively.
The Mutation in katG Ser315Thr observed in isoniazid resistant isolates. The drug susceptibility profile of these isolates were showed resistant to in isoniazid and rifampicin whereas 3 isolates were resistant to Ethambutol drug and Streptomycin respectively alternatively other Seven (7) patients were showing katG gene mutated strain in Ethambutol and Streptomycin resistant cases.
Discussion
We have observed out of 10 samples only one sample shows 132bp region results occurrence of Ser315Thr mutation while the other 9 samples shows 153bp region results no mutation. This suggests these remaining nine patients did not harbor drug resistance. The patients are put on regular treatment regimen except the one patient for multi drug resistance which follows standard treatment for MDR resistance and later follow up of the drug resistance indicates improved in his disease. PCR-RFLP based method for detection of drug resistance is rapid, easy to perform & interpret as compared to other techniques. Therefore this study was undertaken for rapid detection of Ser315Thr katG mutation in Isoniazid resistance population by PCR-RFLP assay. For quick and rapid identification of isoniazid many approaches were reported including real time PCR, single-strand-conformation polymorphism analysis and multiplex-allele-specific PCR but the RFLP approach has several advantages over others methods because it is very cheap, robust, and simple to both perform and interpret, requires PCR equipment [8]. Early detection of resistance to drugs will helpful for treatment modalities and effective control measures to reduce the risk of further disease progression and transmission. The katG and inhA genes mutations account for up to 80% in resistance to isoniazid drug, however majority of mutations are associated with katG gene at specific codon at 315 an estimate of 30–90% of isoniazid-resistant isolates which spans across the world [3,9,10].
Conclusion
Isoniazid along with Rifampicin (RFP) is used as a combination chemotherapy course for tuberculosis (TB) treatment. Mutation at the specific codon position 315 of katG gene is most prevalent method to detect isoniazid resistant Mycobacterium tuberculosis, it will help to screen in high burden countries and in area with higher TB incidence. The technique is feasible for screening TB burden for drug resistance by RFLP method compared to other methods like Genexpert and Line probe assay where the assay require expertise, time consuming ,expensive reagents and expensive instruments.
Polymerase Chain Reaction Restriction Fragment Length Polymorphism (PCR-RFLP) is a reliable and effective tool for the identification of the specific gene alteration in this specific region. The Ser315Thr katG mutation will be a reliable marker for the detection of Isoniazid resistance in M. tuberculosis. The research will also emphases on high prevalence of Ser315Thr mutation in katG in the study population helps in predicting MDR-TB by rapid RFLP Method.
- World Health Organization (2017) Global Tuberculosis 2017 Report. Link: https://goo.gl/u5U6P6
- Ramaswamy S, Musser JM (1998) Molecular genetic basis of antimicrobial agent resistance in Mycobacterium tuberculosis: 1998 update. Tuber Lung Dis 79: 3–29. Link: https://goo.gl/rBqgQ2
- Marahatta SB, Gautam S, Dhital S, Pote N, Jha AK, et al.(2011) kat G (SER 315 THR) Gene Mutation in isoniazid resistant Mycobacterium tuberculosis. kat hmandu Univ Med J 33: 19- 23. Link: https://goo.gl/xrTuV7
- Mokrousov I, Narvskaya O, Otten T, Limeschenko E, Steklova L, et al. (2002) High prevalence of kat G Ser315Thr substitution among isoniazid‐resistant Mycobacterium tuberculosis clinical isolates from northwestern Russia, 1996 to 2001. Antimicrob Agents Chemother 46: 1417‐1424. Link: https://goo.gl/f3duwo
- Rattan A, Kalia A, Ahmad N (1998) Multidrug‐resistant Mycobacterium tuberculosis: molecular perspectives. Emerg Infect Dis 4: 195‐209. Link: https://goo.gl/dgkuqc
- Unissa AN, Selvakumar N, Narayanan S, Suganthi C, Hanna LE (2015) Investigation of Ser315 substitutions within kat G gene in isoniazid-resistant clinical isolates of Mycobacterium tuberculosis from south India. Biomed Res Int 2015: 257983. Link: https://goo.gl/iJoENb
- Shrestha, Raunak (2010) Analysis of kat G Ser315Thr Mutation in Multidrug Resistant Mycobacterium tuberculosis and SLC11A1 Polymorphism in Multidrug Resistance Tuberculosis in Central Development Region of Nepal Using PCR-RFLP Technique: A Pilot Study. Nepal Journal of Biotechnology 1: 14-21. Link: https://goo.gl/cGcqu5
- Caws M, Tho DQ, Duy PM, Lan NT, Hoa DV, et al. (2007) PCR-restriction fragment length polymorphism for rapid, low-cost identification of isoniazid-resistant Mycobacterium tuberculosis. J Clin Microbiol 45: 1789-1793. Link: https://goo.gl/NiNpAx
- Isakova, Jainagul (2018) Mutations of rpoB , kat G, inhA and ahp Genes in Rifampicin and Isoniazid-Resistant Mycobacterium Tuberculosis in Kyrgyz Republic. BMC Microbiology 18: 22. Link: https://goo.gl/7j7Pj3
- Ravibalan T, Samrot AV, Maruthai K, Kommoju V, Kesavan S et al. (2015) Evaluation of Multiplex Polymerase Chain Reaction Assay for the Detection of kat G (S315T) gene Mutation in Mycobacterium tuberculosis Isolates from Puducherr kat y, South India. 9: 2339-2345. Link: https://goo.gl/4z2ocp
Article Alerts
Subscribe to our articles alerts and stay tuned.
 This work is licensed under a Creative Commons Attribution 4.0 International License.
This work is licensed under a Creative Commons Attribution 4.0 International License.


 Save to Mendeley
Save to Mendeley
