Global Journal of Infectious Diseases and Clinical Research
In-house validation of a lamp kit for diagnosis of Plasmodium, Plasmodium falciparum and Plasmodium vivax in Vietnam
National Institute of Malariology, Parasitology and Entomology, Hanoi, Vietnam
Author and article information
Cite this as
Hong Ngoc NT, Huong Binh NT, Hong NV, Thang ND, Huong NT, et al. (2020) In-house validation of a lamp kit for diagnosis of Plasmodium, Plasmodium falciparum and Plasmodium vivax in Vietnam. Glob J Infect Dis Clin Res. 2020; 6(1): 48-53. Available from: 10.17352/2455-5363.000035
Copyright License
© 2020 Hong Ngoc NT, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.Vietnam announced the elimination of malaria in 25 provinces in 2019 and and advance to eliminate malaria nationwide by 2030. Quick, accurate diagnosis and prompt treatment play a very important role in malaria eradication strategies. In this study, we developed the LAMP technique to detect Plasmodium spp., P. falciparum, P. vivax from different kinds of sample such as whole blood, dried blood spot, etc. The advantages of this method included high specificity, fast detection time, and simple equipment use. Primer set was designed for founding on 18S rRNA gene. A positive reaction was visualised with the naked eye, using the color indicator, Malachite Green.
Up to day, Malaria is still considered as a public health problem in tropical and subtropical regions. In 2018, there were approximately 228 million estimated cases of malaria identified globally [1]. According to a report of the national malaria prevention program, in 2019 Vietnam had 4,665 malaria parasites detected by cyanoscopy reduced 3.08% compared to 2018. Rate of parasite / 1,000 people was 0.048. The number of malaria cases gradually decreased through 10-year period from 2010 to 2019. In 2019 malaria patients decreased by 89.16% compared to 2010. Malaria parasites concentrated mainly in the Central and Central Highland provinces. The number of malaria parasites may be lower than they actually are, due to the presence of asymptomatic malaria and the low density of parasites below the detection threshold of a microscope; According to some unpublished reports in Vietnam, this rate can reach 30-50% [2]. Currently, there are three main groups of methods applied in the diagnosis of malaria parasites: Giemsa stained blood smear, rapid diagnostic test (based on immune method), Molecular biology techniques like PCR, Real time-PCR [3]. Giemsa-stained smear test is still the gold standard in the confirmed diagnosis of malaria case [4]. This method is widely used, most popular in many countries around the world, with an average detection threshold of about 50 - 100 parasites/µL of blood from all Plasmodium spp, low test costs, performed even in laboratory conditions and field testing [1]. However, this method is difficult to detect cases with low parasite density, the testing staff must trained, must be retrained regularly, especially in areas with low prevalence of malaria or eliminate areas where there is little opportunity for testing [5-7]. Rapid diagnostic tests (RDTs) are also widely used, especially in areas where microscopy sites have not yet been available, but RDTs still have limitations such as low parasite detection threshold, false positive rate, and long positive time [5,7,8]. The common disadvantage of these two methods is the low threshold for detection of malaria parasites [9,10]. Therefore, it is estimated that a large number of people with low-density parasite infection are not detected. According to WHO (2014), people carrying low-density parasites below detection threshold of GSB and RDTs may be responsible for contributing 20-50% of the pathogen to malaria vector and vice versa [10]. Molecular biology techniques based on DNA analysis have proven to be more sensitive and specific than microscopic detection or rapid test (RDTs) [4]. In particular, traditional PCR techniques such as Nested-PCR, Seminested Multiplex-PCR based on the target gene of 18S -rRNA are widely applied in the diagnostic studies of malaria for high sensitivity and specificity [11-13]. The detection was determined to be between 1-5 parasites /µl of blood [13]. The Real time-PCR technique has a high sensitivity at the LOD > 0.5 KST /µl of blood [14]. However, current DNA amplification techniques are time-consuming, complex and not field-friendly.
To eliminate malaria globally, strategic measures are needed to increase the ability to detect sources of disease. The technical limitations of current testing methods, which requires the development of a highly sensitive diagnostic kit which has a detection threshold comparable to that of DNA amplification techniques in the laboratory, but requires no expensive equipment, faster testing times, is easy for training and can be done in the field. The LAMP technique is a molecular biology technique that uses heat to amplify single strands of DNA, first described by Notomi, et al. [15]. The method has a high amplification efficiency in 15-60 minutes [15]. And then Han ET, et al. [16], developed LAMP to detection of four Plasmodium species for clinical diagnosis [16]. Imai Kazuo, et al. (2017) reported the novel diagnostic method for malaria using LAMP and MinION™ nanopore sequencer [17]. Jaymin C, et al. (2013) developed the RealAmp technique, a LAMP technique combined with real-time isotherms to quickly detect malaria parasites. This study also built a primer pair to detect P. vivax by RealAmp method [18]. Compared with other molecular methods such as PCR, real time PCR ..., LAMP has the advantage of having similar accuracy, simple implementation, especially short execution time and can detect results by naked eye [11,19-22]. Therefore, LAMP is often used to create quick diagnostic kits which are widely popular in the world. LAMP has been utilized to detect malaria parasites mainly using the conventional 18S ribosomal RNA (18S rRNA) gene as the target sequence [6,12,16,23-27]. In this study, we have developed and evaluated the LAMP technique for the detection of Plasmodium genus, Plasmodium falciparum and Plasmodium vivax from whole blood and dry blood spot.
Methods
Samples
All of samples used for positive control purposes were Whole Blood (WB) and Dry Blood Spot (DBS) from human infected with Plasmodium falciparum, Plasmodium vivax, Plasmodium malariae, Plasmodium ovale, and Plasmodium knowlesi. The samples were collected from human Gia Lai province, Vietnam. There were confirmed from WHO EQA program. There used DNA from Plasmodium as positive control and DNA from other parasites was also extracted to determine the specificity.
The use of the human and the collection of samples were approved by the bioethics committee of National Institute of Malariology, Parasitology and Entomology (NIMPE).
DNA extraction protocol
Using the QIAamp DNA microkit of Qiagen (Germany) extracted total DNA of Plasmodium species from WB and DBS samples. We followed the extraction procedure is performed according to the manufacturer’s instructions.
LAMP primer design
Primers used for the LAMP assay were designed based on a highly conserved region of Plasmodium genus, Plasmodium falciparum, Plasmodium vivax genome. Genbank sequences (http://blast.ncbi.nlm.nih.gov/Blast.cgi) (Table 1) including the 18S rRNA gene were tested in silico through BLAST searches and alignment analysis using the Clutal W module of MEGA 7 software, 521 base pairs, 437bp and 562 bp consensus sequences were seclected for the design of specific primers for Plasmodium genus, Plasmodium falciparum, Plasmodium vivax using Primer Explore V5 (http://primerexplorer.jp/lampv5e/index.html).
Optimization of the assay
PCR assay: The outer primers of the LAMP primer designed in this study (the F3 and B3 primers) were tested for their specificity in PCR assay. DNA from WB and DBS which comprised of P. falciparum, P. vivax, P. malariae, P. ovale, P. knowlesi were used as the DNA templates for the specificity test. A standard PCR, using 0.5μM of primers F3 and B3 and 2 μL DNA, was carried out in the total volume of 20μL using HotstarTaq Plus Master Mix kit (Qiagen, Germany). The annealing temperature is 60oC for 30 seconds. PCR products were analysed by electrophoresis in 2% agarose TBE. All reactions were carried out in triplicate.
LAMP assay: There was set up testing different temperatures, MgSO4 concentrations, reaction time, visually detected dye concentrations. LAMP reaction mixtures (25μL) contained 1x Isothermal Amplification buffer (New England Biolabs, UK), MgSO4 (4, 6 or 8 mM) (New England Biolabs, UK), 1.4mM of each dNTPs (Qiagen, Germany), 5pmol of each F3 and B3 primers, 40pmol of each FIB and BIP primers, 20pmol of each LF and LB primers, and 8U of Bst 2.0 DAN polymerase (New England Biolabs, UK) with 5μL of DNA. Different temperatures were tested using a thermocycler (PCR gradient Nesux GX1, Eppendorf) set from 56oC to 65oC for 40 and 60 minutes and then 80oC for 5 minutes. Amplifications were visualy detected by adding Green Malachite at 0.012%, 0.008%, 0.004% and 0.001%. Light blue was observed in successful LAMP reactions and colourless in negative reactions. LAMP reaction products were also analyzed by electrophoresis in 2% TBE agarose. All experiments were completed in all three times.
Preparation of Plasmodium genus, P. falciparum, P. vivax plasmid DNA templates
For the detection and evaluation limits of estmate sensitivity, plasmids containing target regions of the rRNA 18S gene were established for each species and calculated the copy number. The limit of detection (LOD) of LAMP assay was determined by using a ten-fold serial dilution of recombinant plasmid encompassing the target region of the 18S rRNA gene were constructed for all Plasmodium species, P. falciparum and P. vivax (from 10-6 ng/µl to 10-11 ng/µL). The final dilutions were calculated following the formula as mentioned bellow for the corresponding copy number of the 18S rRNA gene as a single copy gene.
The specific PCR primers for cloning were designed based on a highly specific region of the 18S rRNA genes of Plasmodium genus, P. falciparum, P.vivax. Then, genomic DNA was subsequently amplified by PCR, which produced the product 521, 437, 562 bp in size corresponding to Plasmodium genus, P. falciparum, P. vivax. After that, the PCR product was purified and cloned into pUC19 vector (Invitrogen, USA). Concentrations of recombinant plasmid DNA were measured with a Nanophotometer (IMPLEN, Germany), and corresponding copy numbers were calculated as mole multiplies of Avogadro’s number using an online program (http://cels.uri.edu/gsc/cndna.html), following the formula as the number of copies by amount × 6.022 × 1023/length × 109 × 650.
Clinical sample
A total of 200 DBS samples were collected from Gia Lai provinces, Vietnam (between 2017-2018), All these clinical samples were evaluated by LAMP assay comparing with qPCR as gold standard test. The percentage of sensitivity, specificity was calculated in format of 2×2 cross-tabulation Table.
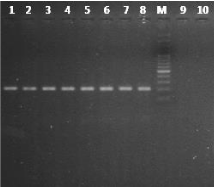
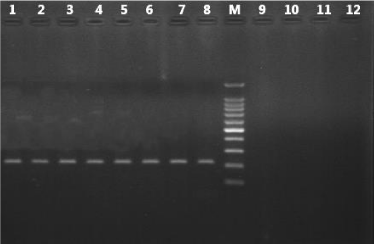
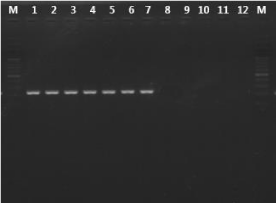
In silico comparisions of LAMP primer sets showed a homology of 100% with Plasmodium genus, P. falciparum and P. vivax. The PCR specificity test using F3 and B3 primers of LAMP Primer amplified DNA from P. vivax, P. falciparum, P. malariae, P. ovale, and P. knowlesi with an amplicon length from 196 to 199bp for Plasmodium genus, 221bp for P. falciparum and 217bp for P. vivax (visualized by gel electrophoresis and UV detection). With P. falciparum and P. vivax no cross reactivity was observed with any of the tested simian malaria parasite species or with the malaria-negative human DNA control (Figure 1) Table 2.
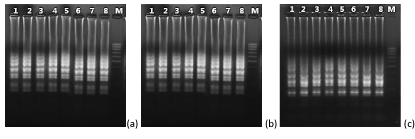
The data revealed that the reaction temperatures at 60oC (Plasmodium genus) and 63oC (P. falciparum and P. vivax) and the concentration of MgSO4 8mM had highest LAMP amplification efficiency (Figure 2).
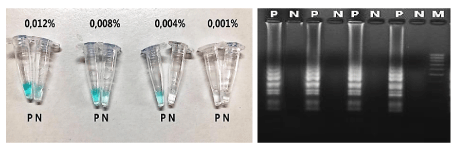
The optimum concentration of MG dye indicated that 0.004% MG was suitable for discrimination of the results as light blue and colourless in positive and negative reactions, respectively (Figure 3).
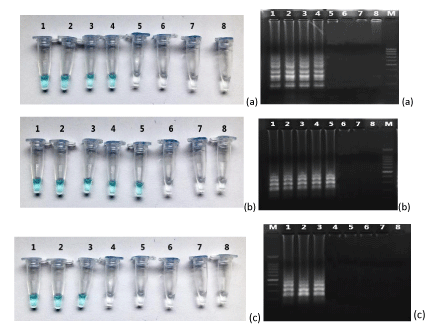
To determine the lower detection limit, 10-fold serial dilutions of each plasmid DNA were amplified. The detection limit for LAMP was 2,97 x 10-2 copies for P. falciparum and 2,85 copies for P. vivax, respectively. The amplification products were visualised on an agarose gel as a ladder of multiple bands. (Figure 4).
Although microscopic examination by Giemsa microscopy has been identified as the gold standard for the diagnosis of malaria parasites. But the detection threshold of this technique is from 20 to 100 parasites/1µl of blood; much higher than the detection threshold of LAMP technique which is about 2-3 parasites/µl of blood; If we compare these two techniques directly, the sensitivity will be very low such as Table 3.
We therefore compared the results of the LAMP test with the qPCR technique, which has similar sensitivity and specificity Tables 4-6.
Total 95 positive samples and 105 negative samples base on qPCR were used. Of the 95 positives, 45 were positive for P. falciparum, 43 were P. vivax positive and seven were positive for remaining Plasmodium spp. The sensitivity was 97.89%, 97.78% and 97.67% for PAN-LAMP, Pf-LAMP and Pv-LAMP assay, respectively. The Specific was 97.14%, 98.06% and 97.45% for PAN-LAMP, Pf-LAMP and Pv-LAMP assay, respectively.
In 2019, Vietnam announced the elimination of malaria in 25 provinces, including 16 provinces in the north, 1 province in the central and 8 provinces in the south. Vietnam’s goal is to be recognized as a country eliminating malaria by 2030 [28]. To achieve this goal, the National Malaria Control Program of Vietnam has been implementing many appropriate strategies such as strengthening monitoring, detection, diagnosis and timely treatment. Every year in Viet Nam, the health system conducts tests of more than 2.2 million blood smears, nearly 500,000 rapid diagnostic tests for malaria [28].The disadvantage of these two techniques is the low detection threshold. Therefore, it is estimated that a large number of people with low-density parasites are not detected. In areas where seasonal malaria is transmitted, people infected with low-density parasites below the detection threshold of blood smear and RDTs are both a regular source of transmission, but also a source of parasite storage through a low infectious season to a high infectious season [28]. Interventions with investigation and treatment to reducing malaria transmission have limited to rely on the detection of antigens using RDTs to identify asymptomatic cases of malaria patients.
Therefore, developing a test with high sensitivity, specificity and easy to do in the field is essential. Numerous studies have demonstrated that the LAMP technique is a highly specific, sensitive, short reaction time and allows the discovery of visual amplification products simply by observing turbidity, fluorescent dye or pH indicator dye. Herein, we have developed and evaluated the LAMP technique for the detection of Plasmodium genus, Plasmodium falciparum and Plasmodium vivax from whole blood and dry blood spot based on the 18S rRNA gene.
We tested our assay specificity with all 5 species P. falciparum, P. vivax, P. malariae, P. ovale, P. knowlesi with satisfying results. However, our assay did not test another nonhuman malaria species. The nucleotide sequence analysis using the BLAST showed that no nonhuman malaria species were related to P. falciparum and P. vivax primer sequences in this study (data not shown).
LAMP assay was carried out for 60 minutes and observed with the naked eye based on the color change in the tubes after amplification. In this study, we used MG as a color indicator dye. The color change of Malachite Green (cationic form) depends on the pH of the solution (pH <2: yellow, pH = 3-9: blue, pH> 10: colorless). Absorption wavelength for MG is 621 nm. In the LAMP-MG assay, positive and negative samples that are easily distinguished by the naked eye with light blue and colorless, respectively. Using of MG as pH-sensitive indicator dye for visual end-point assessment of LAMP products in various infections caused by bacteria [29], protozoa [13,15], and malaria [31] have been reported. Adding MG to LAMP buffer before conducting the reaction did not affect the activity of Bst DNA polymerase, while eliminating the risk of contamination between samples. The optimum concentration of MG used throughout this study was at 0.004%, making positive and negative perfectly differentiated. The higher the MG concentration leads to an increase in false positive, the lower the MG concentration will not be able to distinguish between positive and negative samples. The blue color in the positive tubes still remains in color after the reaction for up to 6 weeks, at room temperature. Time after 6 weeks was not investigated in this study.
Recently, Loopamp™ MALARIA Pan/Pf Detection Kit (Eiken Chemical) has become commercially available for the diagnosis of malaria by the LAMP method. This LAMP kit has good specificity and a lower detection limit of 25 parasites/μL [13,31-34]. The detection limit of our LAMP methods for all Plasmodium species, P. falciparum and P. vivax was 2.89 × 10-1, 2.97 × 10-2 and 2.85 copies/reaction, respectively (5μL of sample DNA could be loaded per reaction). Therefore, the theoretical detection limit of the LAMP assay was equivalent to that of the previously described Loopamp™ MALARIA Pan/Pf and P.v Detection Kit.
The sensitivity and specificity of LAMP assay were evaluated with 200 clinical samples and used qPCR as a gold standard. The results show that the sensitivity and specificity of all assay are over 97%, proving that the LAMP assay in this study has high sensitivity and specificity such as PCR and qPCR. However, PCR and real-time PCR methods require expensive equipments such as thermal cyclers or real-time PCR systems for accurate temperature control, expensive cost and difficult to apply in the field.
Conclusion
LAMP is a molecular method with high sensitivity and specificity. LAMP products are visible to the naked eye after the addition of the MG color indicator, which allows LAMP to be easily deployed in the field in medical facilities.
This study was supported by grants from National key scientific and technological project KC10 / 10.16-20.
- Moody A (2002) Rapid Diagnostic Tests for Malaria Parasites. Clin Microbiol Rev 15: 66-78. Link: https://bit.ly/2P1Zth0
- (2020) National Malaria Control Progamme 2019 report. 80.
- WHO (2013) Proposal for an Evidence Review Group (ERG) on diagnosis of Plasmodium falciparum in low transmission areas. Briefing paper for the Malaria Policy Advisory Committee. Link: https://bit.ly/3g6oiEz
- WHO (2015) World Malaria Report, Geneva, World Health Organization. Link: https://bit.ly/3f5JKrT
- McMorrow M, Aidoo M, Kachur S (2011) Malaria rapid diagnostic tests in elimination settings—can they find the last parasite? Clin Microbiol Infect 17: 1624-1631. Link: https://bit.ly/332vzBC
- Mohon AN, Getie S, Jahan N, Alam MS, Pillai DR (2019) Ultrasensitive loop mediated isothermal amplification (US-LAMP) to detect malaria for elimination. Malar J 18: 350. Link: https://bit.ly/2P2Zb9O
- Mohon AN, Lee LDY, Bayih AG, Folefoc A, Guelig D, et al. (2016) NINALAMP compared to microscopy, RDT, and nested PCR for the detection of imported malaria. Diagn Microbiol Infect Dis 85: 149-153. Link https://bit.ly/2WXpl1Z
- Wongsrichanalai C, Barcus MJ, Muth S, Sutamihardja A, Wernsdorfer WH (2007) A review of malaria diagnostic tools: microscopy and rapid diagnostic test (RDT). Am J Trop Med Hyg 77: 119–127. Link: https://bit.ly/2DeiGJ y
- Mens P, Spieker N, Omar S, Heijnen M, Schallig H, et al. (2007) Is molecular biology the best alternative for diagnosis of malaria to microscopy? ADN comparison between microscopy, antigen detection and molecular tests in rural Kenya and urban Tanzania. Trop Med Int Health 12: 238–244. Link: https://bit.ly/2WZlhOv
- WHO/HTM/GMP/2014.7 (2014) Policy brief on malaria diagnostics in low-transmission setting. Link: https://bit.ly/306DiwE
- Milne L, Kyi M, Chiodini P, Warhurst D (1994) Accuracy of routine laboratory diagnosis of malaria in the United Kingdom. J Clin Pathol 47: 740-742. Link: https://bit.ly/39yZ5jk
- Poschl B, Waneesorn J, Thekisoe O, Chutipongvivate S, Karanis P (2010) Comparative diagnosis of malaria infections by microscopy, nested PCR, and LAMP in northern Thailand. Am J Trop Med Hyg 83: 56–60. Link: https://bit.ly/39D2Bt 5
- Snounou G, Viriyakosola S, Zhua XP, Jarraa W, Pinheirob L, et al. (1993) High sensitivity of detection of human malaria parasites by the use of nested olymerase chain reaction. Mol Biochem Parasitol 61: 315-320. Link: https://bit.ly/30WjJWX
- Wu L, Lotus LH, Hannah S, Walker PGT, Ghani AZC, et al. (2015) Coparison of diganostics for dectection of asymtomatic Plasmodium falciparum infections to inform control and elimination strategies. Nature 528: S86-93. Link: https://bit.ly/2P3rzbK
- Notomi T, Okayama H, Masubuchi H, Yonekawa T, Watanabe K, et al. (2000) Loop-mediated isothermal amplification of DNA. Nucleic Acids Res 28: E63. Link: https://bit.ly/2Da1Prq
- Han ET, Watanabe R, Sattabongkot J, Khuntirat B, Sirichaisinthop J, et al. (2007) Detection of four Plasmodium species by genus- and species-specific loopmediated isothermal amplification for clinical diagnosis. J Clin Microbiol 45: 2521–2528. Link: https://bit.ly/2X2KUOE
- Hopkins H, González IJ, Polley SD, Angutoko P, Ategeka J, et al. (2013) Highly sensitive detection of malaria parasitemia in a malaria-endemic setting: performance of a new loop-mediated isothermal amplification kit in a remote clinic in Uganda. J Infect Dis 208: 645-652. Link: : https://bit.ly/2EsbUAL
- Patel JC, Oberstaller J, Xayavong M, Narayanan J, DeBarry JD, et al. (2013) Real-Time Loop-Mediated Isothermal Amplification (RealAmp) for the Species-Specific Identification of Plasmodium vivax. Plos one J 8: e54986. Link: https://bit.ly/30RuTfz
- Lucchi NW, Gaye M, Diallo MA, Goldman IF, Ljolje D, et al. (2016) Evaluation of the illumigene malaria LAMP: a robust molecular diagnostic tool for malaria parasites. Sci Rep 6: 36808. Link: https://bit.ly/3jNRLVO
- Perera RS, Ding XC, Tully F, Oliver J, Bright N, et al. (2017) Development and clinical performance of high throughput loop-mediated isothermal amplification for detection of malaria. PLoS One12: e0171126. Link: https://bit.ly/3f6YTt5
- Polley SD, González IJ, Mohamed D, Daly R, Bowers K, et al. (2013) Clinical evaluation of a loop-mediated amplification kit for diagnosis of imported malaria. J Infect Dis 208: 637-644. Link: https://bit.ly/30ankSd
- Rypien C, Chow B, Chan W, Church D, Pillai DR (2017) Detection of Plasmodium spp. infection by the illumigene malaria assay compared to reference microscopy and real-time PCR. J Clin Microbiol 55: 3037-3045. Link: https://bit.ly/332sRM7
- Chen JH, Lu F, Lim CS, Kim JY, Ahn HJ, et al. (2010) Detection of Plasmodium vivax infection in the Republic of Korea by loop-mediated isothermal amplification (LAMP). Acta Trop 113: 61–65. Link: https://bit.ly/3g7Xh3 g
- Imai K, Tarumoto N, Misawa K, Runtuwene LR, Sakai J, et al. (2017) A novel diagnostic method for malaria using loop-mediated isothermal amplification (LAMP) and MinION™ nanopore sequencer. B BMC Infect Dis 17: 621. Link: Link: https://bit.ly/39z4JC5
- Ling LY, Lai MY, Fong MY, Jelip J, Mahmud R (2016) Loop-Mediated Isothermal Amplification Assay for Identification of Five Human Plasmodium Species in Malaysia. Am J Trop Med Hyg 94: 336-339. Link: https://bit.ly/2X0gVXK
- Paris DH, Imwong M, Faiz AM, Hasan M, Yunus EB, et al. (2007) Loopmediated isothermal PCR (LAMP) for the diagnosis of falciparum malaria. Am J Trop Med Hyg 77: 972–976. Link: https://bit.ly/3hGltdC
- Poon LL, Wong BW, Ma EH, Chan KH, Chow LM, et al. (2006) Sensitive and inexpensive molecular test for falciparum malaria: detecting Plasmodium falciparum DNA directly from heat-treated blood by loop-mediated isothermal amplification. Clin Chem 52: 303–306. Link: https://bit.ly/3jMrxD9
- WHO (2018) National Malaria Programme Review – VietNam. Link: https://bit.ly/3hNVKjD
- Farnia P, Masjedi MR, Mohammadi F, Tabarsei P, Farnia P, et al (2008) Colorimetric detection of multidrugresistant or extensively drug-resistant tuberculosis by use of malachite green indicator dye. J Clin Microbiol 46: 796–799. Link: https://bit.ly/3f3XHGz
- Lucchi NW, Ljolje D, Silva-Flannery L, Udhayakumar V (2016) Use of malachite green-loop mediated isothermal amplification for detection of Plasmodium spp. parasites. PLoS One 11: e0151437. Link: https://bit.ly/3hMBD5j
- Falade CO, Ajayi IO, Nsungwa-Sabiiti J, Siribié M, Diarra A, et al. (2016) Malaria rapid diagnostic tests and malaria microscopy for guiding malaria treatment of uncomplicated fevers in Nigeria and prereferral cases in 3 African countries. Clin Infect Dis 63: S290-297. Link: https://bit.ly/3hIFwbe
- Nzelu CO, Cáceres AG, Guerrero-Quincho S, TineoVillafuerte E, Rodriquez-Delfin L, et al. (2016) A rapid molecular diagnosis of cutaneous leishmaniasis by colorimetric malachite green-loop-mediated isothermal amplification (LAMP) combined with an FTA card as a direct sampling tool. Acta Trop 153: 116-119. Link: https://bit.ly/2CTNjnV
- Sriworarat C, Phumee A, Mungthin M, Leelayoova S, Siriyasatien P (2015) Development of loop-mediated isothermal amplification (LAMP) for simple detection of Leishmania infection. Parasit Vectors 8: 591. Link: https://bit.ly/302RMNM
- WHO (2019) World Malaria Report, Geneva, World Health Organization. Link: https://bit.ly/3hIE7l7
Article Alerts
Subscribe to our articles alerts and stay tuned.
 This work is licensed under a Creative Commons Attribution 4.0 International License.
This work is licensed under a Creative Commons Attribution 4.0 International License.


 Save to Mendeley
Save to Mendeley
